Stata and R in a jupyter notebook¶
The jupyter notebook project is now designed to be a ‘language agnostic’ web-application front-end for any one of many possible software language kernels. We’ve been mostly using python but there are in fact several dozen other language kernels that can be made to work with it including Julia, R, Matlab, C, Go, Fortran and Stata.
The ecosystem of libraries and packages for scientific computing with python is huge and constantly growing but there are still many statistics and econometrics applications that are available as built-in or user-written modules in Stata that have not yet been ported to python or are just simply easier to use in Stata. On the other hand there are some libraries such as python pandas and different visualization libraries such as seaborn or matplotlib that give features that are not available in Stata.
Fortunately you don’t have to choose between using Stata or python, you can use them both together, to get the best of both worlds.
Jupyter running an R kernel¶
R is a powerful open source software environment for statistical computing. R has R markdown which allows you to create R-markdown notebooks similar in concept to jupyter notebooks. But you can also run R inside a jupyter notebook (indeed the name ‘Jupyter’ is from Julia, iPython and R).
See my notebook with notes on Research Discontinuity Design for an example of a jupyter notebook running R. To install an R kernel see the IRkernel project.
Jupyter with a Stata Kernel¶
Kyle Barron has created a stata_kernel that offers several useful features including code-autocompletion, inline graphics, and generally fast responses.
For this to work you must have a working licensed copy of Stata version 14 or greater on your machine.
Python and Stata combined in the same notebook¶
Sometimes it may be useful to combine python and Stata in the same
notebook. Ties de Kok has written a nice python library called
ipystata that allows one to
execute Stata code in codeblocks inside an ipython notebook when
preceded by a %%stata magic command.
This workflow allows you to pass data between python and Stata sessions and to display Stata plots inline. Compared to the stata_kernel option the response times are not quite as fast.
The remainder of this notebook illustrates the use of ipystata.
A sample ipystata session¶
For more details see the example notebook and documentation on the ipystata repository.
In [1]:
%matplotlib inline
import seaborn as sns
import pandas as pd
import statsmodels.formula.api as smf
import ipystata
The following opens a Stata session where we load a dataset and
summarize the data. The -o flag following the `%%Stata``` magic
instructs it to output or return the dataset in Stata memory as a pandas
dataframe in python.
In [2]:
%%stata -o life_df
sysuse lifeexp.dta
summarize
(Life expectancy, 1998)
Variable | Obs Mean Std. Dev. Min Max
-------------+---------------------------------------------------------
region | 68 1.5 .7431277 1 3
country | 0
popgrowth | 68 .9720588 .9311918 -.5 3
lexp | 68 72.27941 4.715315 54 79
gnppc | 63 8674.857 10634.68 370 39980
-------------+---------------------------------------------------------
safewater | 40 76.1 17.89112 28 100
Let’s confirm the data was returned as a pandas dataframe:
In [3]:
life_df.head(3)
Out[3]:
| region | country | popgrowth | lexp | gnppc | safewater | |
|---|---|---|---|---|---|---|
| 0 | Eur & C.Asia | Albania | 1.2 | 72 | 810.0 | 76.0 |
| 1 | Eur & C.Asia | Armenia | 1.1 | 74 | 460.0 | NaN |
| 2 | Eur & C.Asia | Austria | 0.4 | 79 | 26830.0 | NaN |
A simple generate variable command and ols regression in Stata:
In [4]:
%%stata -o life_df
gen lngnppc = ln(gnppc)
regress lexp lngnppc
(5 missing values generated)
Source | SS df MS Number of obs = 63
-------------+---------------------------------- F(1, 61) = 97.09
Model | 873.264865 1 873.264865 Prob > F = 0.0000
Residual | 548.671643 61 8.99461709 R-squared = 0.6141
-------------+---------------------------------- Adj R-squared = 0.6078
Total | 1421.93651 62 22.9344598 Root MSE = 2.9991
------------------------------------------------------------------------------
lexp | Coef. Std. Err. t P>|t| [95% Conf. Interval]
-------------+----------------------------------------------------------------
lngnppc | 2.768349 .2809566 9.85 0.000 2.206542 3.330157
_cons | 49.41502 2.348494 21.04 0.000 44.71892 54.11113
------------------------------------------------------------------------------
And the same regression using statsmodels and pandas:
In [5]:
model = smf.ols(formula = 'lexp ~ lngnppc',
data = life_df)
results = model.fit()
print(results.summary())
OLS Regression Results
==============================================================================
Dep. Variable: lexp R-squared: 0.614
Model: OLS Adj. R-squared: 0.608
Method: Least Squares F-statistic: 97.09
Date: Tue, 08 May 2018 Prob (F-statistic): 3.13e-14
Time: 14:25:48 Log-Likelihood: -157.57
No. Observations: 63 AIC: 319.1
Df Residuals: 61 BIC: 323.4
Df Model: 1
Covariance Type: nonrobust
==============================================================================
coef std err t P>|t| [0.025 0.975]
------------------------------------------------------------------------------
Intercept 49.4150 2.348 21.041 0.000 44.719 54.111
lngnppc 2.7683 0.281 9.853 0.000 2.207 3.330
==============================================================================
Omnibus: 21.505 Durbin-Watson: 1.518
Prob(Omnibus): 0.000 Jarque-Bera (JB): 46.140
Skew: -1.038 Prob(JB): 9.57e-11
Kurtosis: 6.643 Cond. No. 52.7
==============================================================================
Warnings:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
Back to ipython¶
Let’s change one of the variables in the dataframe in python:
In [6]:
life_df.popgrowth = life_df.popgrowth * 100
In [7]:
life_df.popgrowth.mean()
Out[7]:
97.20587921142578
And now let’s push the modified dataframe into the Stata dataset with
the -d flag:
In [8]:
%%stata -d life_df
summarize
Variable | Obs Mean Std. Dev. Min Max
-------------+---------------------------------------------------------
index | 68 33.5 19.77372 0 67
region | 68 .5 .7431277 0 2
country | 0
popgrowth | 68 97.20588 93.11918 -50 300
lexp | 68 72.27941 4.715315 54 79
-------------+---------------------------------------------------------
gnppc | 63 8674.857 10634.68 370 39980
safewater | 40 76.1 17.89112 28 100
lngnppc | 63 8.250023 1.355677 5.913503 10.59613
A Stata plot:
In [9]:
%%stata -d life_df --graph
graph twoway (scatter lexp lngnppc) (lfit lexp lngnppc)
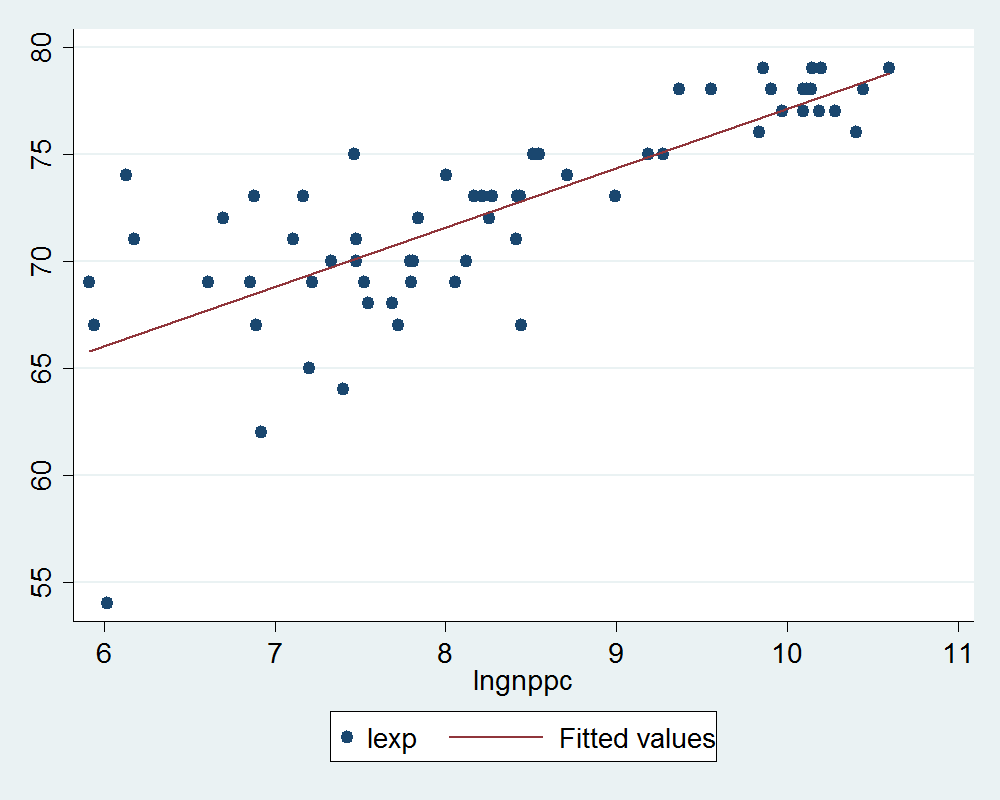
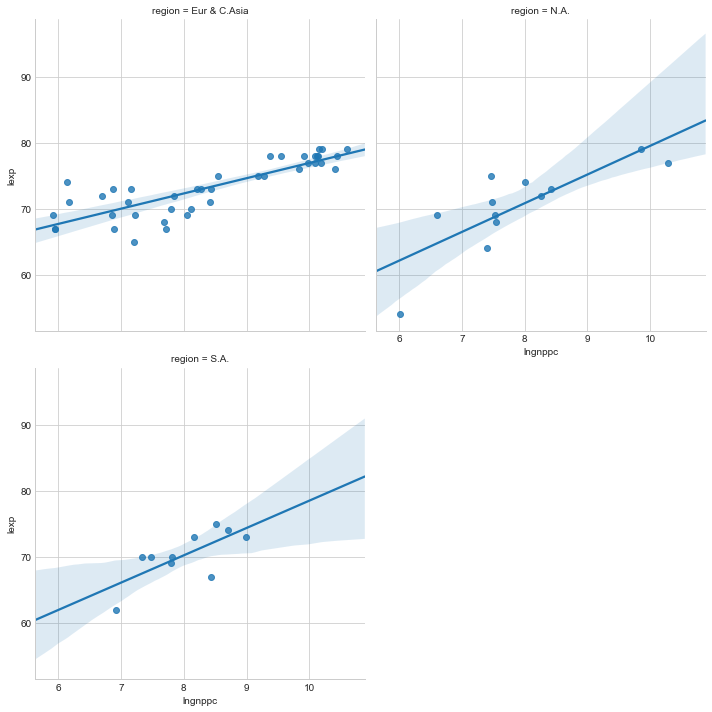
Now on the python side use lmplot from the seaborn library to graph a similar scatter and fitted line but by region.
In [10]:
sns.set_style("whitegrid")
g=sns.lmplot(y='lexp', x='lngnppc', col='region', data=life_df,col_wrap=2)